paired end sequencing wikipedia
The library prep kits that it uses are optimized for a variety of applications including targeted gene small genome and amplicon sequencing 16S. The similarities between PE and MP reads include.
Get 1 month free of our Silver Membership including 2 additional DNA reports.
. Ad Gene Expression Profiling Chromosome Counting Epigenetic Changes Molecular Analysis. Illumina sequencing by synthesis technology supports both single-read and paired-end libraries. Requires the same amount of DNA as single-read genomic DNA or cDNA sequencing.
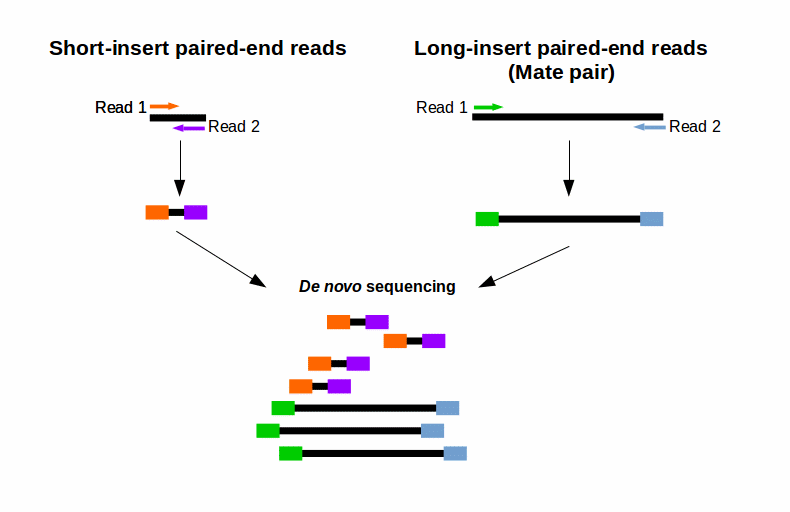
Combining data from mate pair sequencing with those from short-insert paired-end reads provides increased information for maximising sequencing coverage across a genome 1. Simple workflow allows generation of unique ranges of insert sizes. In almost all cases short paired-end reads trump single-end reads that sequence approximately the same number of bases.
Simple workflow allows generation of unique ranges of insert sizes. It is capable of automated paired-end reads and up to 15 Gb per run delivering over 600 bases of sequence data per read. In genetics shotgun sequencing is a method used for sequencing random DNA strands.
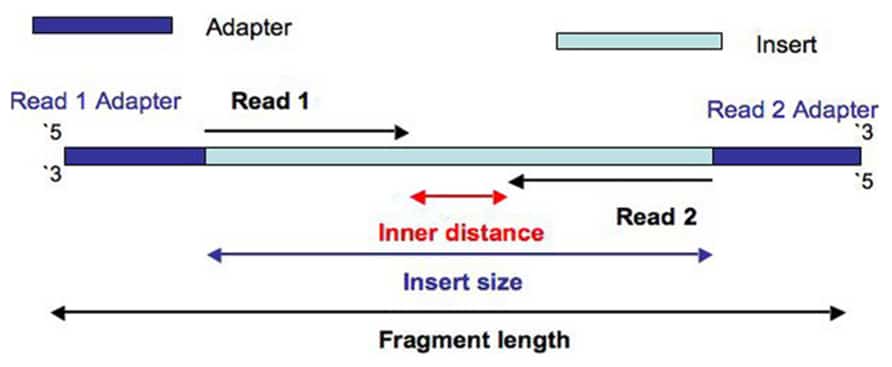
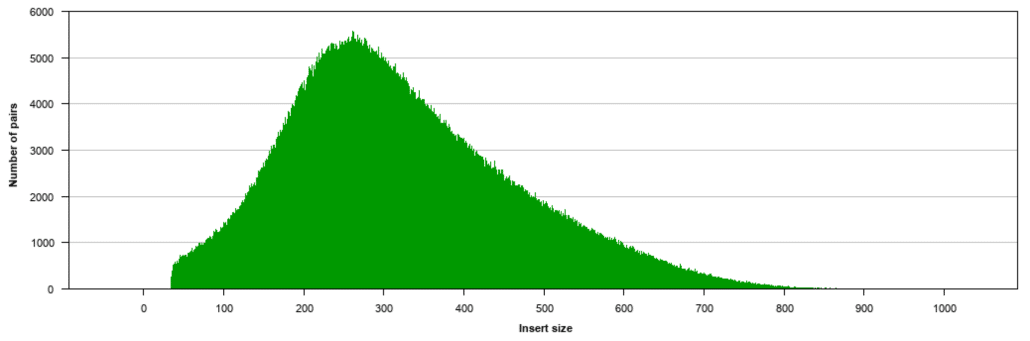
Now lets get started. Sequenziare in una o due direzioni. The inner mate distance between the two reads is 200 bp creating an insert size of 400 bp.
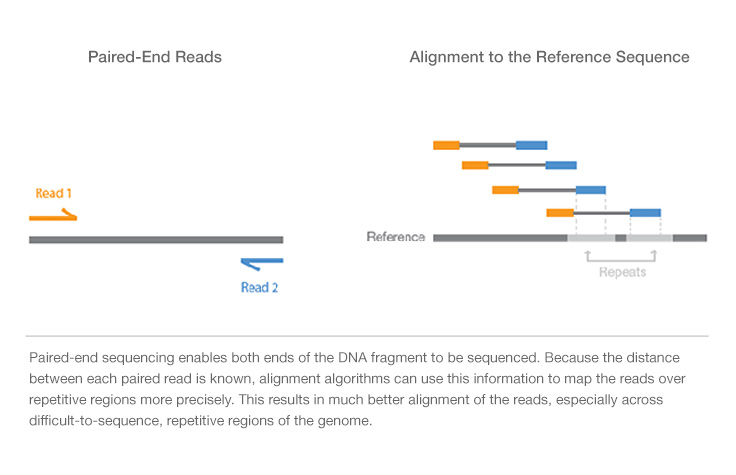
Paired-end reading improves the ability to identify the relative positions of various reads in the genome making it much more effective than single-end reading in resolving structural rearrangements such as. The reads were then mapped back to the reference using BWA aln and sampe. Mate pair sequencing involves generating long-insert paired-end DNA libraries useful for a number of sequencing applications including.
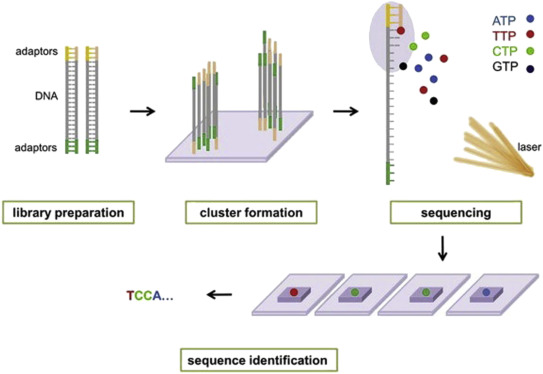
For your De novo genome assembly Fig. We do not sell or share your data. A sequence identifier with information about the sequencing run and the cluster.
Does not require methylation of DNA or restriction digestion. Can be used for bisulfite sequencing. The enzyme tagmentase fragments the.
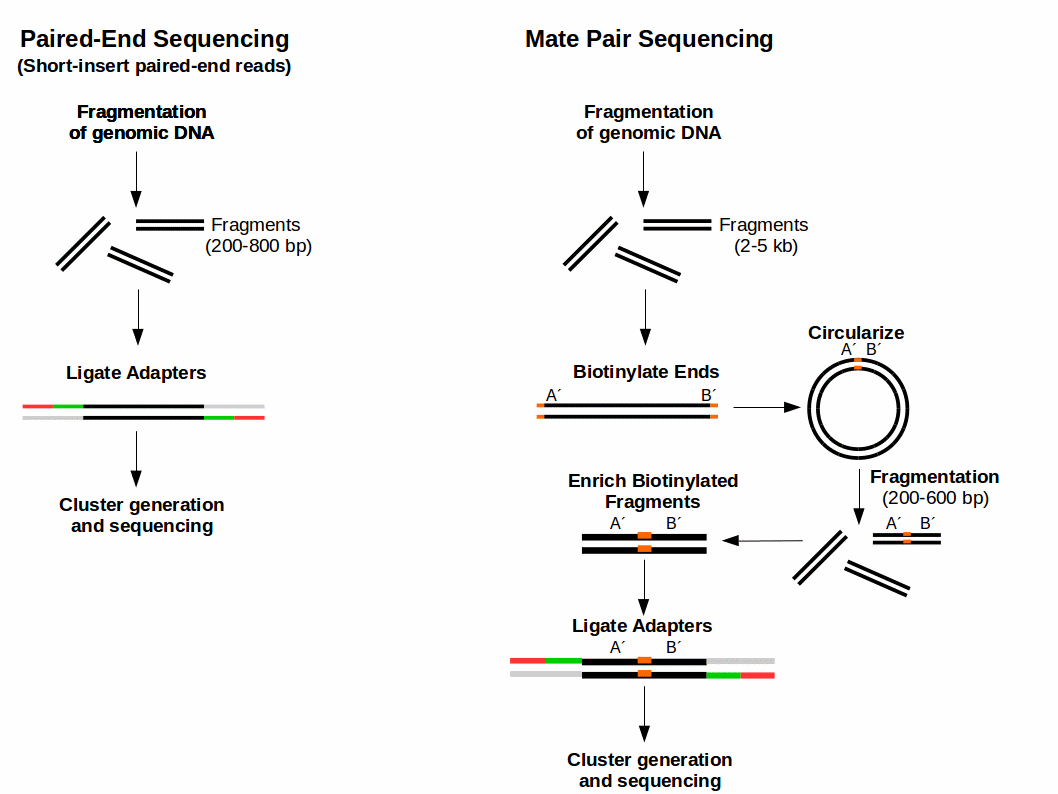
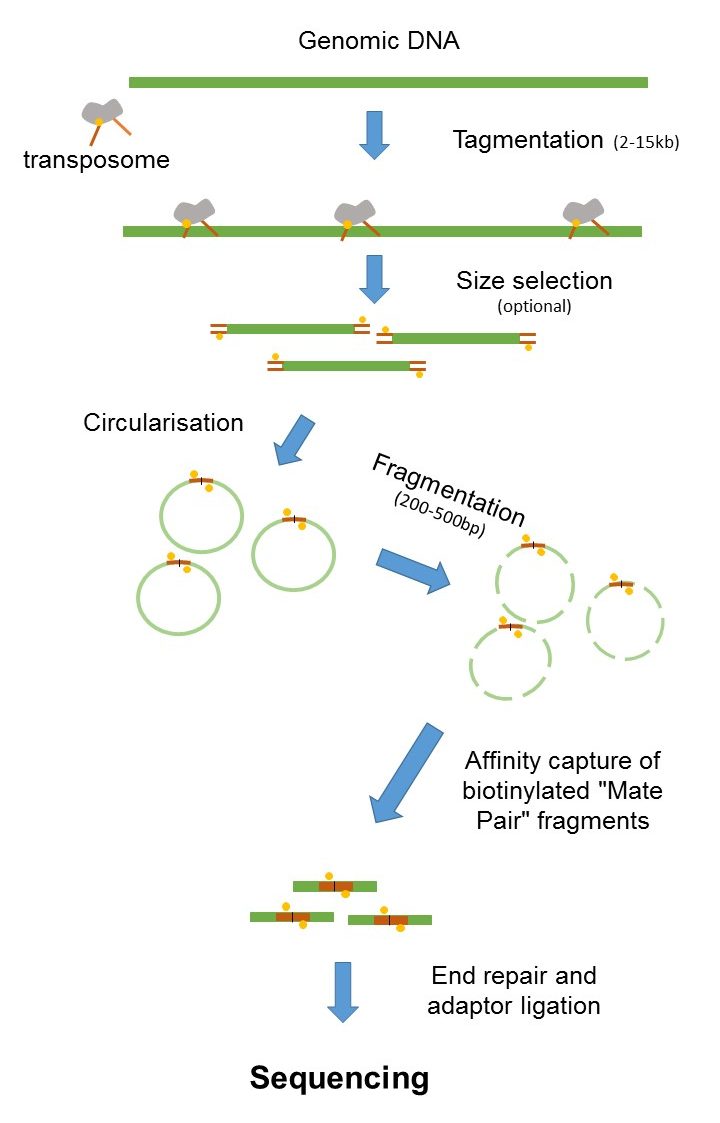
The HiSeq 2500 System has been discontinued. The figure shows the workflow for mate-pair library preparation for Illumina sequencing. Short paired-end reads provide an additional benefit in that they can identify a large fraction of differentially expressed isoforms for the cost of gene-level analysis with longer single-end reads.
Due modi di fare NGS Come già visto in Cosa è una read una read è una corta sequenza di DNA di sintesi che viene prodotta durante la reazione di sequenziamento. Sequencing depth or library size. Broad Range of Applications.
It is named by analogy with the rapidly expanding quasi-random shot grouping of a shotgun. Learn about the difference between Paired-End and Single-Run sequencing and why the former creates more precise alignments than the latter especiall. Can be used for bisulfite sequencing.
I frammenti che si utilizzano nella costruzione della. For each cluster that passes filter a single sequence is written to the corresponding samples R1 FASTQ file and for a paired-end run a single sequence is also written to the samples R2 FASTQ file. Combining data generated from mate pair library sequencing with that from short-insert paired-end reads provides a powerful combination of read lengths for maximal sequencing coverage across the genome.
Ad Privacy first so your data belongs to you. Reads come in pairs. The MiSeq System facilitates your research with a wide range of sequencing applications.
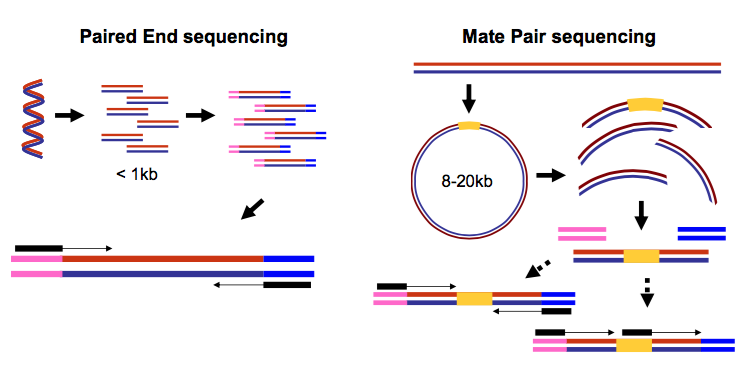
SBS technology offers a short-insert paired-end capability for high-resolution genome sequencing as well as long-insert paired-end reads for efficient sequence assembly de novo sequencing and more. Paired-end sequencing means sequencing both ends of the cDNA fragments and aligning the forward and reverse reads as read pairs Figure 8. We intend to continue providing full support of the instrument and supplying the reagents through February 28th 2023.
Each entry in a FASTQ files consists of 4 lines. For paired-end RNA-Seq use the following kits with an alternate fragmentation protocol followed by standard Illumina paired-end cluster generation and sequencing. Configure the system to sequence a trio in one day or up to 48 genomes in 2 days for the most comprehensive coverage.
For the first test I took some sequence from the human genome hg19 and created two 100 bp reads from this region. The larger inserts mate pairs can pair reads across greater distances. Fast and Accurate Next-Generation Sequencing Results Enabled by Ion Torrent Technology.
Does not require methylation of DNA or restriction digestion. While the underlying principles between PE and MP reads have strong similarities there are inherent differences that are crucial to understand. Paired end runs read from one end to the other end and then start another round of reading from the opposite end.
Il sequenziamento NGS può essere fatto utilizzando single-end reads o paired-end reads. Sign in and order today. 4 before high-throughput sequencing the quality of the library should be verified using sanger sequencing wherein the long sequencing.
In DNA sequencing lingo the words paired-end PE and mate-pair MP are frequently used interchangeably. Requires the same amount of DNA as single-read genomic DNA or cDNA sequencing. We offer several instruments that support the same applications as the HiSeq 2500 System.
Therefore they are able to better cover highly. The preparation of mate pair libraries is designed to allow classical paired-end sequencing of both ends of a fragment with an original size of several kilobases. Broad Range of Applications.
The chain-termination method of DNA sequencing Sanger sequencing can only be used for short DNA strands of 100 to 1000 base pairsDue to this size limit longer sequences are subdivided. Since the beginning of 2013 this preparation has been based on Nextera technology. The combination of short inserts and longer reads increases the ability to fully.
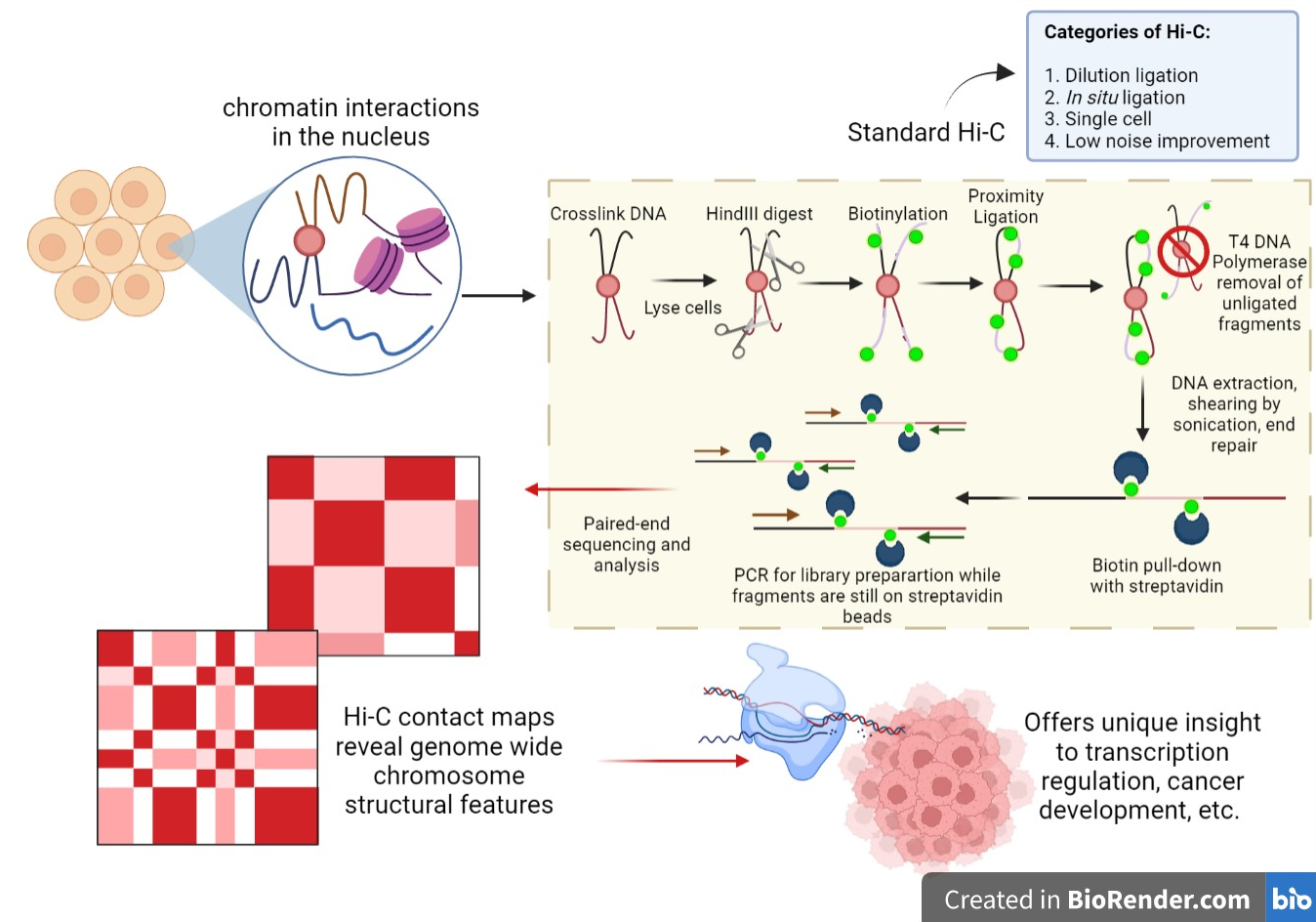
Single read runs are faster cheaper and are typically sufficient for profiling or counting studies such as RNA-Seq or ChIP-Seq. We recommend using our Sequencing Platform Comparison Tool to find the best instrument for. Any platform that can allow for the ligated fragments to be sequenced across the nhei junction roche 454 or by paired-end or mate-paired reads illumina ga and hiseq platforms would be suitable for hi-c.
With single read runs the sequencing instrument reads from one end of a fragment to the other end. This can be very helpful e. In paired-end reading it starts at one read finishes this direction at the specified read length and then starts another round of reading from the opposite end of the fragment.
Paired-end reads are preferable for de novo transcript discovery or isoforms expression analysis as well as to characterise poorly annotated transcriptomes. Paired end sequencing refers to the fact that the fragment s sequenced were sequenced from both ends and not just the one as was true for first. Ad Complete your research with 300000 products.

Bacterial Chromosome Replication
What Is Mate Pair Sequencing For
What Is Mate Pair Sequencing For
Hi C Genomic Analysis Technique Wikipedia

Sequencing Introduction To Sequencing By Synthesis Sbs Youtube
What Are Paired End Reads The Sequencing Center
Difference Between Whole Genome Sequencing And Next Generation Sequencing Difference Between
Mate Pair Sequencing France Genomique

What Are Paired End Reads The Sequencing Center
Manipulating Ngs Data With Galaxy Galaxy Community Hub

Bacterial Chromosome Replication

Fundamental Principles In Cardiovascular Genetics Sciencedirect